Blueprint for the affordable genome
The following is part two in a series of stories that highlight recent discoveries enabled by the Stampede supercomputer. Read part one, three, four and five to find out how Stampede is making a difference through science and engineering.
Aleksei Aksimentiev, a professor of physics at the University of Illinois-Urbana Champaign, used the National Science Foundation-supported Stampede supercomputer to explore a cutting-edge method of DNA sequencing. The method uses an electric field to drive a strand of DNA through a small hole, or "nanopore," either in silicon or a biological membrane.
By controlling this process precisely and measuring the change in ionic current as the DNA strands move through the pore of the membrane, the sequencer can read each base pair in order.
"Stampede is by far the best computer system my group has used over the past 10 years," Aksimentiev said. "Being able to routinely obtain 40-80 nanoseconds of molecular dynamic simulations in 24 hours, regardless of the systems' size, has been essential for us to make progress with rapidly evolving projects."
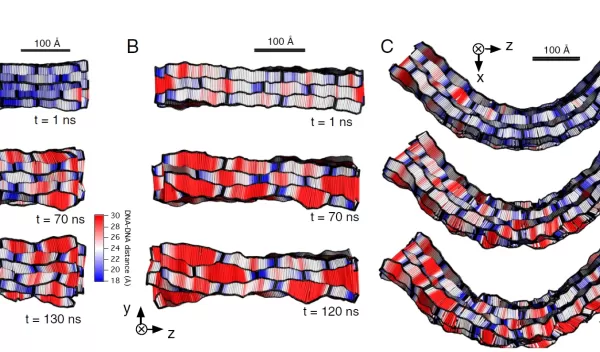
Aksimentiev and his group showed that localized heating can be used to stretch DNA, which significantly increases the accuracy of nanopore DNA sequencing. In addition, he and his team used an all-atom molecular dynamics method to accurately describe DNA origami objects, making it possible to engineer materials for future applications in biosensing, drug delivery and nano-electronics. These results were published in ACS Nano and the Proceedings of the National Academy of Sciences.


