News Release 04-008
Protein Data Bank Opens New Era With Broader Support
Nearly 24,000 molecules and growing, accessible collection advances biology
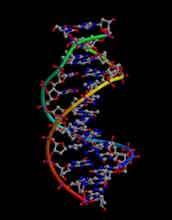
The structure of DNA in its most common configuration (B-form).
January 21, 2004
This material is available primarily for archival purposes. Telephone numbers or other contact information may be out of date; please see current contact information at media contacts.
ARLINGTON, Va.—The assets of the Protein Data Bank (PDB) just keep growing.
The PDB holds the three-dimensional structures of nearly 24,000 proteins and other macromolecules in its growing – and publicly accessible – collection. Its holdings profile DNAs, RNAs, viruses, and various proteins, such as enzymes central to photosynthesis, growth, development and brain function.
This month, with a doubling in the number of the federal agencies supporting it, the PDB begins a new five-year, $30 million management era, the National Science Foundation announced today. The chapter opens following a new international agreement announced last month to pool and coordinate the deposit of molecular structure data globally.
Mary Clutter, assistant director for NSF's Directorate for Biological Sciences, said, "The Protein Data Bank is a treasure chest of shared discoveries. This new agreement will ensure that it continues to serve biologists around the world as its collection grows and diversifies.
"Biological processes involve small molecular machines," she said. "Understanding how these machines function often begins with knowing how their parts are structured, how they fit together. Thus, to have these molecular structures archived comprehensively, centrally and consistently is of enormous value across the spectrum of biological research, from genomics to systems biology.
"And because of the data bank's openness and accessibility, individual researchers - and humanity as a whole - will continue to benefit from the collective research of thousands of biologists," Clutter said.
For example, the collection includes the intricate membrane-channel proteins recognized in the 2003 Nobel Prize in Chemistry.
The structure of another PDB deposit, the enzyme carbonic anhydrase, also permeates biology. Showcased as the PDB's January 2004 "Molecule of the Month," it is crucial for photosynthesis in plants and bacteria, the building of coral reefs and many fundamental processes in animals – such as bone formation, breathing and muscle contraction.
NSF has supported the Protein Data Bank continuously since 1975. A multi-agency support partnership first formed in 1989. For the past five years, that partnership has included NSF, the National Institute of General Medical Sciences (NIGMS), the Department of Energy (DOE) and the National Library of Medicine (NLM). The partnership has been expanded now to include the National Cancer Institute (NCI), the National Center for Research Resources (NCRR), the National Institute of Biomedical Imaging and Bioengineering (NIBIB), and the National Institute of Neurological Disorders and Stroke (NINDS).
The agreement, which began Jan. 1, calls for the PDB to continue to be managed by the three members of the Research Collaboratory for Structural Bioinformatics (RCSB): Rutgers, The State University of New Jersey; the San Diego Supercomputer Center at the University of California, San Diego; and the University of Maryland/National Institute of Standards and Technology's Center for Advanced Research in Biotechnology.
Last month, the RCSB announced an international partnership to establish a worldwide PDB, coordinating with similar efforts at the Institute for Protein Research at Osaka University in Japan and at the European Bioinformatics Institute (EBI) in the United Kingdom.
The expansion of federal agency partnerships and international participation mirrors the expansion in opportunities for progress in a new era of structure-informed research.
According to James Cassatt of NIGMS, "The use of structures has revolutionized the development of new drugs, including that of all of the HIV protease inhibitors. The use of these drugs as part of combination therapy is prolonging the lives of people infected with HIV."
The PDB collection includes a wide variety of medically important structures, including enzymes and other proteins associated with influenza, HIV, SARS and other viruses; parts of prion proteins (including the bovine form implicated in Mad Cow Disease or BSE); the amyloid peptide associated with Alzheimer's disease; and the p53 tumor-suppressor protein associated with a wide variety of human cancers.
The PDB also serves the Department of Energy's Genomics:GTL program, which explores the biology of microbes to seek new ways to remediate environmental contamination, sequester carbon dioxide and generate energy from biomass. According to Aristides Patrinos, director of the Office of Biological and Environmental Research in DOE's Office of Science, knowing the structures of key molecules will help scientists understand "the protein machines that carry out the many functions of microbial cells in communities."
As the sole international repository for comprehensive structural data of large biological molecules, the PDB serves researchers and educators in academic, industrial and biotechnical pursuits.
When the data bank was first established in 1971, it contained seven structures. After 25 years, that number grew to slightly more than 5,000 structures. Three years later, there were more than 10,000. Deposits keep coming, and their data keeps generating interest worldwide: During 2003, more than 4,600 new molecular structures were added, and, on an average day, bank visitors downloaded various structural files more than 120,000 times.
According to PDB Director Helen Berman, "When the PDB started, it was felt that the data contained in protein structures would provide the information needed to understand the molecular underpinnings for a host of biological processes. This vision is being realized, and it is now even more important that the data be preserved and publicly available from a single source."
The structural data comes from experiments using x-ray crystallography, nuclear magnetic resonance, electron microscopy and other methods. After a scientist submits a structure, the experimental data – the deposit – is validated and annotated. Coordinating with the biological journals that publish the discovery of new protein structures, the PDB also ensures that the data is available in the public domain.
As the PDB grows and evolves, one of its central challenges will be the expanded integration of its wealth of information with other biological data, images and research articles.
According to Kim Henrick of the European Bioinformatics Institute, "The PDB must expand both in the storage and annotation of protein production information and into other 3-D structure fields with linkages made to electron microscopy (EM) data. EM experimental data will make an enormous impact in the next five years in molecular biology."
Over the next five years, the PDB's challenges will also include keeping up with the increasing complexity and volume of deposited structures, meeting the demands for more complex queries, and providing more detailed annotation of the experiments and the structures.
Along with serving scientists, the PDB also serves as an educational resource for students and educators at all levels, thus another challenge is to meet the needs of an expanding, diverse and global user community.
-NSF-
-
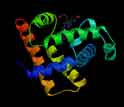
Myoglobin, the first protein structure to be determined at high resolution.
Credit and Larger Version -
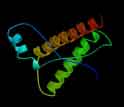
Structure of a portion of the bovine prion protein, a molecule associated with mad cow disease.
Credit and Larger Version -
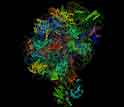
Structure of the largest of the two subunits of the ribosome - the 'protein factory' of the cell.
Credit and Larger Version
Note to editors and news directors:
These related materials are available:
Fact Sheet - Timeline for structural biology, PDB
Sidebar - Examples, Impacts of PDB structures
Additional images are available from www.pdb.org.
Protein Data Bank Senior Project Personnel:
Helen M. Berman (primary contact), (732) 445-4667, berman@rcsb.rutgers.edu
http://rutchem.rutgers.edu/faculty/berman.html
Department of Chemistry and Chemical Biology
Rutgers, The State University of New Jersey
Piscataway, NJ 08854
Philip E. Bourne, (858) 534-8301, bourne@sdsc.edu
http://www.sdsc.edu/~bourne
San Diego Supercomputer Center
University of California, San Diego
San Diego, CA 92093
Judith L. Flippen-Anderson, (732) 445-0103; flippen@rcsb.rutgers.edu
Department of Chemistry and Chemical Biology
Rutgers, The State University of New Jersey
Piscataway, NJ 08854
Gary L. Gilliland, (301) 738-6262; gary.gilliland@nist.gov
University of Maryland Biotechnology Institute
Center for Advanced Research in Biotechnology
National Institute of Standards and Technology
Rockville, MD 20850
John Westbrook, (732) 445-4290; jwest@rcsb.rutgers.edu
Department of Chemistry and Chemical Biology
Rutgers, The State University of New Jersey
Piscataway, NJ 08854
A full contact sheet is available here: http://www.nsf.gov/od/lpa/news/04/fscontacts_pdb_04.htm.
Background resources, related news available on the web:
- Protein Data Bank (PDB) – The single worldwide repository for the processing and distribution of 3-D biological macromolecular structure, it has more than 23,000 structures in its collection.
http://www.rcsb.org/pdb/ - The Research Collaboratory for Structural Bioinformatics (RCSB) – The non-profit consortium that manages the Protein Data Bank, it focuses on advancing the study of the 3-D structure of biological macromolecules to understand better the function of biological systems. It works through joint grants to provide free public resources to further the fields of bioinformatics and biology.
http://www.rcsb.org/index.html - NSF Fact Sheet: Timeline for Structural Biology and the Protein Data Bank
- NSF Fact Sheet: PDB Examples/Impacts from Fundamental Biology to Disease
Related news releases:
RCSB News Release, Dec. 2, 2003: International Collaborators to Form the Worldwide Protein Data Bank - The Research Collaboratory for Structural Bioinformatics (RCSB), the Macromolecular Structure Database at the EMBL-European Bioinformatics Institute (MSD-EBI), and Protein Data Bank Japan (PDBj) have announced a collaboration to form the Worldwide Protein Data Bank (www.wwpdb.org/):
http://www.wwpdb.org/news.html
http://www.rcsb.org/pdb/pdb_news2003.html#wwpdb
Agency links:
National Science Foundation: www.nsf.gov
National Institute of General Medical Sciences (NIGMS): http://www.nigms.nih.gov/
Department of Energy (DOE) Office of Science: www.sc.doe.gov
National Library of Medicine (NLM): http://www.nlm.nih.gov/
National Cancer Institute (NCI): http://www.nci.nih.gov/
National Center for Research Resources (NCRR): http://www.ncrr.nih.gov/
National Institute of Biomedical Imaging and Bioengineering (NIBIB): http://www.nibib1.nih.gov/
National Institute of Neurological Disorders and Stroke (NINDS): http://www.ninds.nih.gov/
Media Contacts
Sean Kearns, NSF, (703) 292-7963, email: skearns@nsf.gov
Program Contacts
Chris L. Greer, NSF, (703) 292-8470, email: cgreer@nsf.gov
The U.S. National Science Foundation propels the nation forward by advancing fundamental research in all fields of science and engineering. NSF supports research and people by providing facilities, instruments and funding to support their ingenuity and sustain the U.S. as a global leader in research and innovation. With a fiscal year 2023 budget of $9.5 billion, NSF funds reach all 50 states through grants to nearly 2,000 colleges, universities and institutions. Each year, NSF receives more than 40,000 competitive proposals and makes about 11,000 new awards. Those awards include support for cooperative research with industry, Arctic and Antarctic research and operations, and U.S. participation in international scientific efforts.
Connect with us online
NSF website: nsf.gov
NSF News: nsf.gov/news
For News Media: nsf.gov/news/newsroom
Statistics: nsf.gov/statistics/
Awards database: nsf.gov/awardsearch/
Follow us on social
Twitter: twitter.com/NSF
Facebook: facebook.com/US.NSF
Instagram: instagram.com/nsfgov