News Release 08-029
All Eyes and Ears on the Corn Genome
NSF-supported researchers complete draft sequence in less than three years

In 2005, NSF, DOE, and USDA funded the sequencing of the corn genome.
February 28, 2008
This material is available primarily for archival purposes. Telephone numbers or other contact information may be out of date; please see current contact information at media contacts.
A consortium of researchers led by the Genome Sequencing Center (GSC) at Washington University in St. Louis, Mo., announced today the completion of a draft sequence of the corn genome.
In the fall of 2005 the NSF, in partnership with the U.S. Department of Agriculture (USDA) and the Department of Energy (DOE), awarded $32 million to two projects to sequence the corn genome. The goal of the project led by the Washington University GSC is to develop a map-based genome sequence for the B73 inbred line of corn.
This groundbreaking sequencing project was funded by the NSF under the auspices of the National Plant Genome Initiative (NPGI). The initiative, which began in 1998, is an ongoing effort to understand the structure and function of all plant genes at levels from the molecular and organismal, to interactions within ecosystems. NPGI's focus is on plants of economic importance and plant processes of potential economic value. Sequencing the corn genome is one of the major goals of the current initiative.
"Corn is one of the most economically important crops for our nation," said NSF Director, Arden L. Bement, Jr. "Completing this draft sequence of the corn genome constitutes a significant scientific advance and will foster growth for the agricultural community and the economy as a whole."
According to the USDA, more than 80 million acres of land in the United States is devoted to growing corn, accounting for more than 90 percent of the total value of feed grain.
"Corn is a vitally important crop," said Rick Wilson, lead investigator and director of the GSC. "Scientists will now be able to accurately and efficiently probe the genome to develop new varieties of corn that increase crop yields and resist drought and disease. The information we glean from the corn genome is also likely to be applicable to other grains, such as rice, wheat and barley."
Sequencing the corn genome has been an immense and daunting task. At 2.5 billion base pairs covering 10 chromosomes, this genome's size is comparable to that of the human genome. Corn also has one of the most complex genomes of any known organism and is one of the most challenging genomes sequenced to date. The draft sequence will allow researchers to begin to uncover the functional components of individual genes as well as develop an overall picture of the genome organization. Completing the draft sequence, which covers about 95 percent of the genome, is an important milestone on the way to refining the complete genome sequence.
"Creating a completed draft of the corn genome brings us one step closer to our goal of understanding the functional genetic components that influence hybrid vigor, drought and pest resistance, and asexual plant reproduction or apomixis - all special traits that make corn valuable," said James Collins, head of the Biological Sciences Directorate at the NSF.
The National Corn Growers Association, a strong supporter of the sequencing project and an advocate of the NPGI, notes that elucidating the complete sequence and structure of all corn genes, associated functional sequences and their locations on corn's genetic and physical map, has many potential benefits. These include: creating a model for other major genome sequencing projects, enhancing the efficiency of modern corn breeding programs, increasing understanding of corn's important agronomic traits, and strengthening the physical and intellectual scientific processes of the genetic research community.
Pam Johnson, chairman of the Research and Business Development Action Team for the National Corn Growers Association, adds, "This effort is especially critical at this time in history, when the growing global population looks to corn and other plants to supply food, feed, bioenergy and biobased materials. It is time to learn the language of corn as a model that has great potential and economic significance."
Collaborators contributing to the GSC corn genome research include: Rod Wing from the University of Arizona; W. Richard McCombie, Robert Martienssen, Doreen Ware, and Lincoln Stein from Cold Springs Harbor Laboratory; Patrick Schnable and Srinivas Aluru from Iowa State University; and Richard Wilson and Sandy Clifton from Washington University.-NSF-
-
Corn ranks among the world's major grain crops and dominates American agriculture.
Credit and Larger Version -
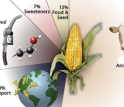
Corn, also known as maize, is used to produce a myriad of products.
Credit and Larger Version -
To sequence the maize genome, scientists first collect and purify DNA from maize plants.
Credit and Larger Version
Media Contacts
Lily Whiteman, NSF, (703) 292-8070, email: lwhitema@nsf.gov
Program Contacts
Jane Silverthorne, NSF, (703) 292-8470, email: jsilvert@nsf.gov
Erin (Liz) Lawrence, NSF, (703) 292-8997, email: elawrenc@nsf.gov
The U.S. National Science Foundation propels the nation forward by advancing fundamental research in all fields of science and engineering. NSF supports research and people by providing facilities, instruments and funding to support their ingenuity and sustain the U.S. as a global leader in research and innovation. With a fiscal year 2023 budget of $9.5 billion, NSF funds reach all 50 states through grants to nearly 2,000 colleges, universities and institutions. Each year, NSF receives more than 40,000 competitive proposals and makes about 11,000 new awards. Those awards include support for cooperative research with industry, Arctic and Antarctic research and operations, and U.S. participation in international scientific efforts.
Connect with us online
NSF website: nsf.gov
NSF News: nsf.gov/news
For News Media: nsf.gov/news/newsroom
Statistics: nsf.gov/statistics/
Awards database: nsf.gov/awardsearch/
Follow us on social
Twitter: twitter.com/NSF
Facebook: facebook.com/US.NSF
Instagram: instagram.com/nsfgov